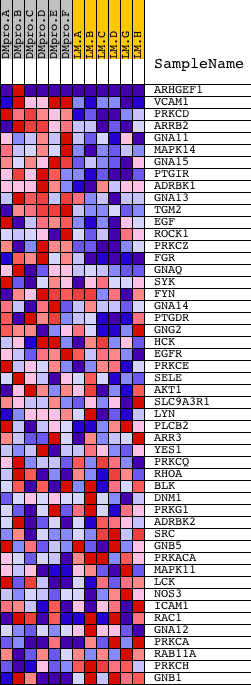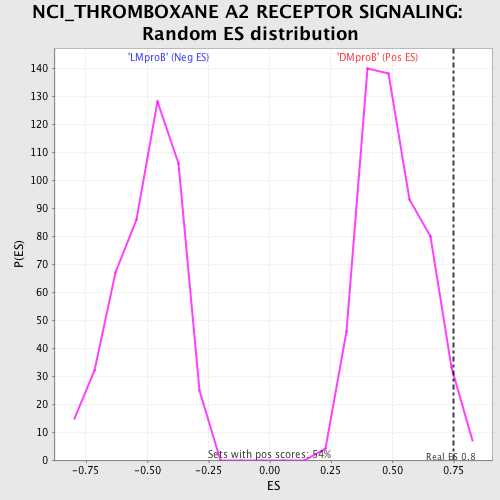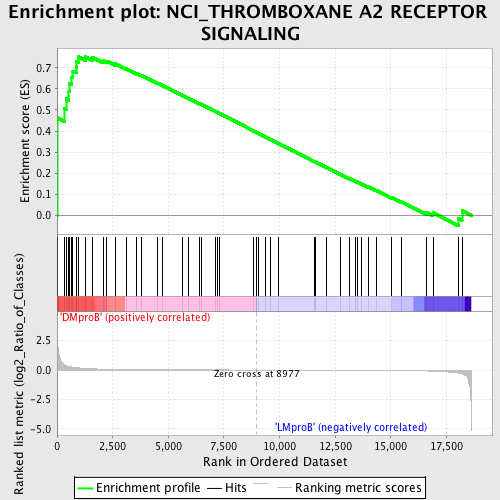
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | Set_04_DMproB_versus_LMproB.phenotype_DMproB_versus_LMproB.cls #DMproB_versus_LMproB |
| Phenotype | phenotype_DMproB_versus_LMproB.cls#DMproB_versus_LMproB |
| Upregulated in class | DMproB |
| GeneSet | NCI_THROMBOXANE A2 RECEPTOR SIGNALING |
| Enrichment Score (ES) | 0.7519021 |
| Normalized Enrichment Score (NES) | 1.4892466 |
| Nominal p-value | 0.03142329 |
| FDR q-value | 0.71476394 |
| FWER p-Value | 1.0 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | ARHGEF1 | 1952 18343 3996 | 7 | 3.371 | 0.4629 | Yes | ||
| 2 | VCAM1 | 5851 | 314 | 0.450 | 0.5082 | Yes | ||
| 3 | PRKCD | 21897 | 406 | 0.366 | 0.5536 | Yes | ||
| 4 | ARRB2 | 20806 | 533 | 0.295 | 0.5873 | Yes | ||
| 5 | GNA11 | 3300 3345 19670 | 547 | 0.290 | 0.6265 | Yes | ||
| 6 | MAPK14 | 23313 | 659 | 0.251 | 0.6549 | Yes | ||
| 7 | GNA15 | 19671 | 712 | 0.234 | 0.6843 | Yes | ||
| 8 | PTGIR | 18376 | 857 | 0.200 | 0.7040 | Yes | ||
| 9 | ADRBK1 | 23960 | 877 | 0.196 | 0.7300 | Yes | ||
| 10 | GNA13 | 20617 | 946 | 0.184 | 0.7516 | Yes | ||
| 11 | TGM2 | 2657 | 1284 | 0.134 | 0.7519 | Yes | ||
| 12 | EGF | 15169 | 1577 | 0.103 | 0.7504 | No | ||
| 13 | ROCK1 | 5386 | 2079 | 0.067 | 0.7326 | No | ||
| 14 | PRKCZ | 5260 | 2237 | 0.059 | 0.7322 | No | ||
| 15 | FGR | 4723 | 2604 | 0.043 | 0.7185 | No | ||
| 16 | GNAQ | 4786 23909 3685 | 3100 | 0.029 | 0.6958 | No | ||
| 17 | SYK | 21636 | 3583 | 0.021 | 0.6728 | No | ||
| 18 | FYN | 3375 3395 20052 | 3774 | 0.019 | 0.6651 | No | ||
| 19 | GNA14 | 23907 | 4505 | 0.012 | 0.6275 | No | ||
| 20 | PTGDR | 21866 | 4736 | 0.011 | 0.6166 | No | ||
| 21 | GNG2 | 4790 | 5629 | 0.007 | 0.5696 | No | ||
| 22 | HCK | 14787 | 5921 | 0.006 | 0.5548 | No | ||
| 23 | EGFR | 1329 20944 | 6418 | 0.005 | 0.5288 | No | ||
| 24 | PRKCE | 9575 | 6472 | 0.005 | 0.5266 | No | ||
| 25 | SELE | 14074 | 7111 | 0.003 | 0.4927 | No | ||
| 26 | AKT1 | 8568 | 7187 | 0.003 | 0.4891 | No | ||
| 27 | SLC9A3R1 | 11250 | 7314 | 0.003 | 0.4827 | No | ||
| 28 | LYN | 16281 | 8848 | 0.000 | 0.4002 | No | ||
| 29 | PLCB2 | 5262 | 8945 | 0.000 | 0.3950 | No | ||
| 30 | ARR3 | 24282 9305 | 8971 | 0.000 | 0.3937 | No | ||
| 31 | YES1 | 5930 | 9040 | -0.000 | 0.3901 | No | ||
| 32 | PRKCQ | 2873 2831 | 9370 | -0.001 | 0.3724 | No | ||
| 33 | RHOA | 8624 4409 4410 | 9583 | -0.001 | 0.3612 | No | ||
| 34 | BLK | 3205 21791 | 9957 | -0.002 | 0.3413 | No | ||
| 35 | DNM1 | 2777 8857 | 11580 | -0.005 | 0.2546 | No | ||
| 36 | PRKG1 | 5289 | 11612 | -0.005 | 0.2536 | No | ||
| 37 | ADRBK2 | 11499 | 12122 | -0.006 | 0.2270 | No | ||
| 38 | SRC | 5507 | 12717 | -0.008 | 0.1960 | No | ||
| 39 | GNB5 | 2985 9029 3153 | 13157 | -0.009 | 0.1736 | No | ||
| 40 | PRKACA | 18549 3844 | 13408 | -0.010 | 0.1616 | No | ||
| 41 | MAPK11 | 2264 9618 22163 | 13481 | -0.011 | 0.1591 | No | ||
| 42 | LCK | 15746 | 13697 | -0.012 | 0.1492 | No | ||
| 43 | NOS3 | 16906 885 | 13976 | -0.014 | 0.1360 | No | ||
| 44 | ICAM1 | 19545 | 14337 | -0.016 | 0.1188 | No | ||
| 45 | RAC1 | 16302 | 15046 | -0.024 | 0.0840 | No | ||
| 46 | GNA12 | 9023 | 15497 | -0.032 | 0.0641 | No | ||
| 47 | PRKCA | 20174 | 16623 | -0.077 | 0.0141 | No | ||
| 48 | RAB11A | 7013 | 16896 | -0.096 | 0.0126 | No | ||
| 49 | PRKCH | 21246 | 18055 | -0.260 | -0.0140 | No | ||
| 50 | GNB1 | 15967 | 18200 | -0.321 | 0.0224 | No |